Note
Go to the end to download the full example code.
Brain segmentation with median_otsu#
We show how to extract brain information and mask from a b0 image using DIPY’s
segment.mask module.
First import the necessary modules:
import matplotlib.pyplot as plt
import numpy as np
from dipy.core.histeq import histeq
from dipy.data import get_fnames
from dipy.io.image import load_nifti, save_nifti
from dipy.segment.mask import bounding_box, crop, median_otsu
Download and read the data for this tutorial.
The scil_b0 dataset contains different data from different companies and
models. For this example, the data comes from a 1.5 Tesla Siemens MRI.
data_fnames = get_fnames(name="scil_b0")
data, affine = load_nifti(data_fnames[1])
data = np.squeeze(data)
Segment the brain using DIPY’s mask module.
median_otsu returns the segmented brain data and a binary mask of the
brain. It is possible to fine tune the parameters of median_otsu
(median_radius and num_pass) if extraction yields incorrect results
but the default parameters work well on most volumes. For this example,
we used 2 as median_radius and 1 as num_pass
In case of holes or minor errors in the mask, add the argument
finalize_mask=True to apply a final step to the mask, which will fix
those issues.
b0_mask, mask = median_otsu(data, median_radius=2, numpass=1)
Saving the segmentation results is very easy. We need the b0_mask, and
the binary mask volumes. The affine matrix which transform the image’s
coordinates to the world coordinates is also needed. Here, we choose to save
both images in float32.
fname = "se_1.5t"
save_nifti(fname + "_binary_mask.nii.gz", mask.astype(np.float32), affine)
save_nifti(fname + "_mask.nii.gz", b0_mask.astype(np.float32), affine)
Quick view of the results middle slice using matplotlib.
sli = data.shape[2] // 2
plt.figure("Brain segmentation")
plt.subplot(1, 2, 1).set_axis_off()
plt.imshow(histeq(data[:, :, sli].astype("float")).T, cmap="gray", origin="lower")
plt.subplot(1, 2, 2).set_axis_off()
plt.imshow(histeq(b0_mask[:, :, sli].astype("float")).T, cmap="gray", origin="lower")
plt.savefig(f"{fname}_median_otsu.png", bbox_inches="tight")
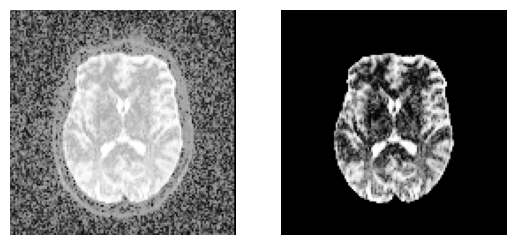
An application of median_otsu for brain segmentation.
We can also crop the outputs to remove the largest possible number of
background voxels. This makes outputted data significantly smaller. We can
use the crop function to crop the data using the bounding box of the
mask.
# 1. Regenerate mask with NEW parameters (radius=4, numpass=4)
b0_mask_crop, mask_crop = median_otsu(data, median_radius=4, numpass=4)
# 2. Calculate bounding box and crop
mins, maxs = bounding_box(mask_crop)
b0_mask_crop = crop(b0_mask_crop, mins, maxs)
mask_crop = crop(mask_crop, mins, maxs)
Saving cropped data as demonstrated previously.
save_nifti(fname + "_binary_mask_crop.nii.gz", mask_crop.astype(np.float32), affine)
save_nifti(fname + "_mask_crop.nii.gz", b0_mask_crop.astype(np.float32), affine)
Total running time of the script: (0 minutes 19.899 seconds)