Visualization#
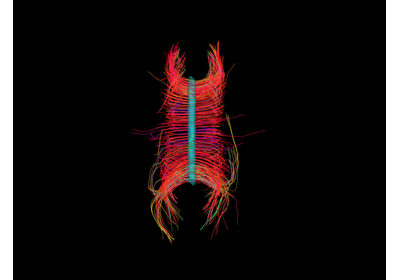
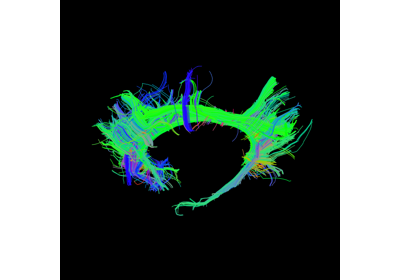
Visualization of ROI Surface Rendered with Streamlines
Visualization of ROI Surface Rendered with Streamlines
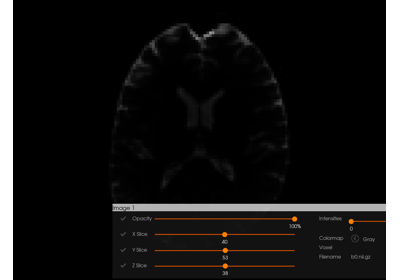
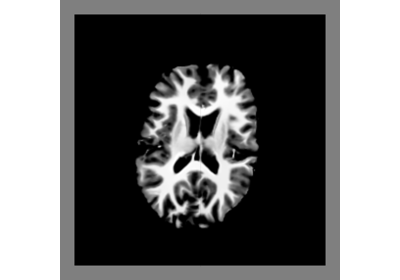
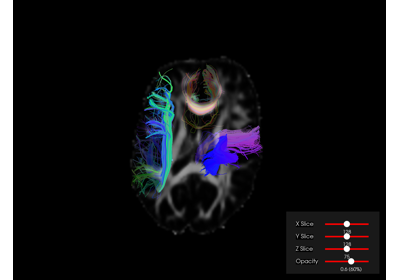
Interactive Visualization with DIPY Horizon (Python)
Interactive Visualization with DIPY Horizon (Python)
Site Navigation
News
Help
Section Navigation

Visualization of ROI Surface Rendered with Streamlines

Interactive Visualization with DIPY Horizon (Python)